Discover Toolkit
With the DISCOVER module, teams can combine their knowledge and data with AI algorithms built to understand biology — leading to new, high performance bio based products faster than ever before. Our AI models allow you to converge on an optimal product ten times faster than using traditional approaches.
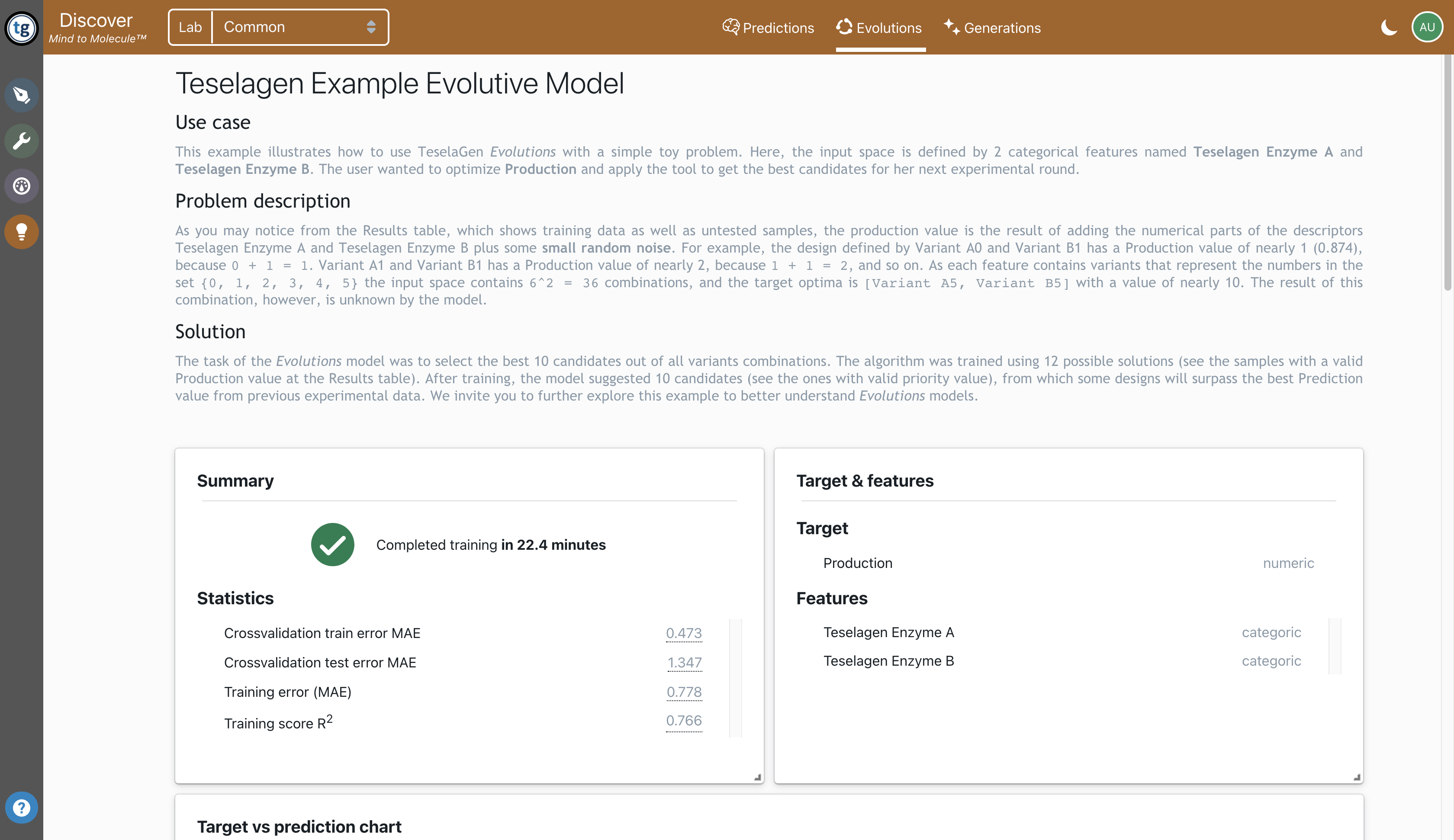
Train predictive models
We implement predictive models that can be used to predict target values based of quantitative or qualitative inputs. These predictive models can be used to predict the phenotype of different biological products, such as the binding affinity of an antibody or the titer of a metabolite. Biological and chemical entities can be highly dimensional but can be mapped to an embedding space. We implement different embeddings to represent DNA molecules, proteins, and chemical compounds, to efficiently train predictive models.
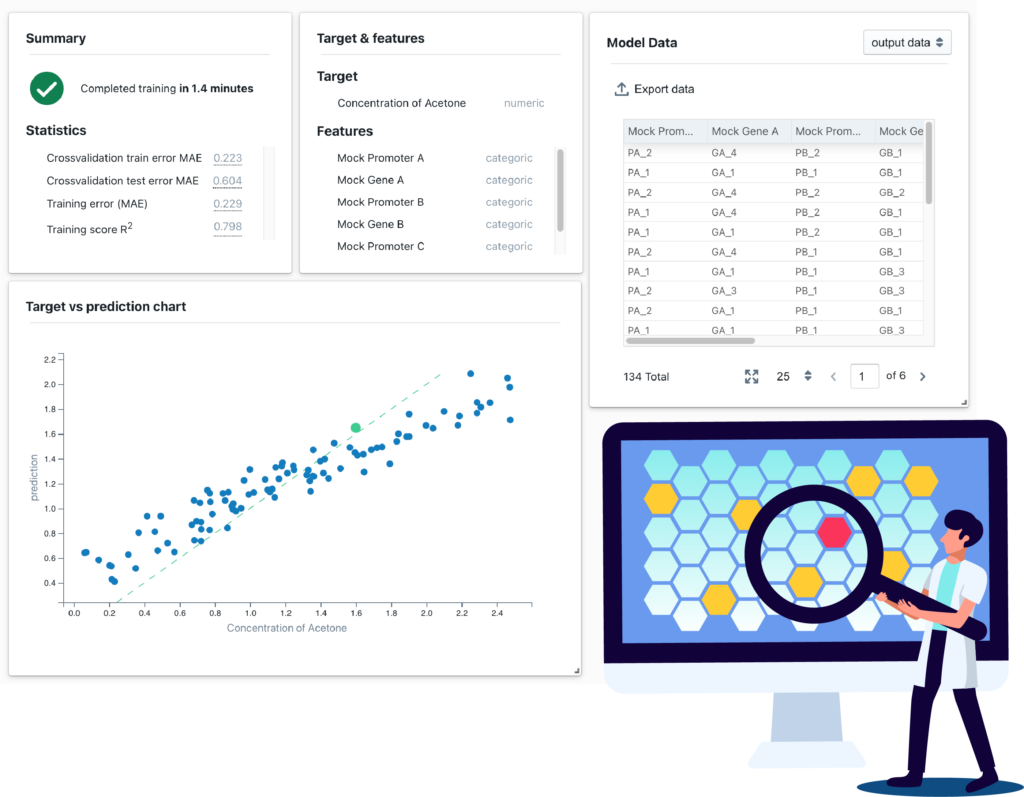
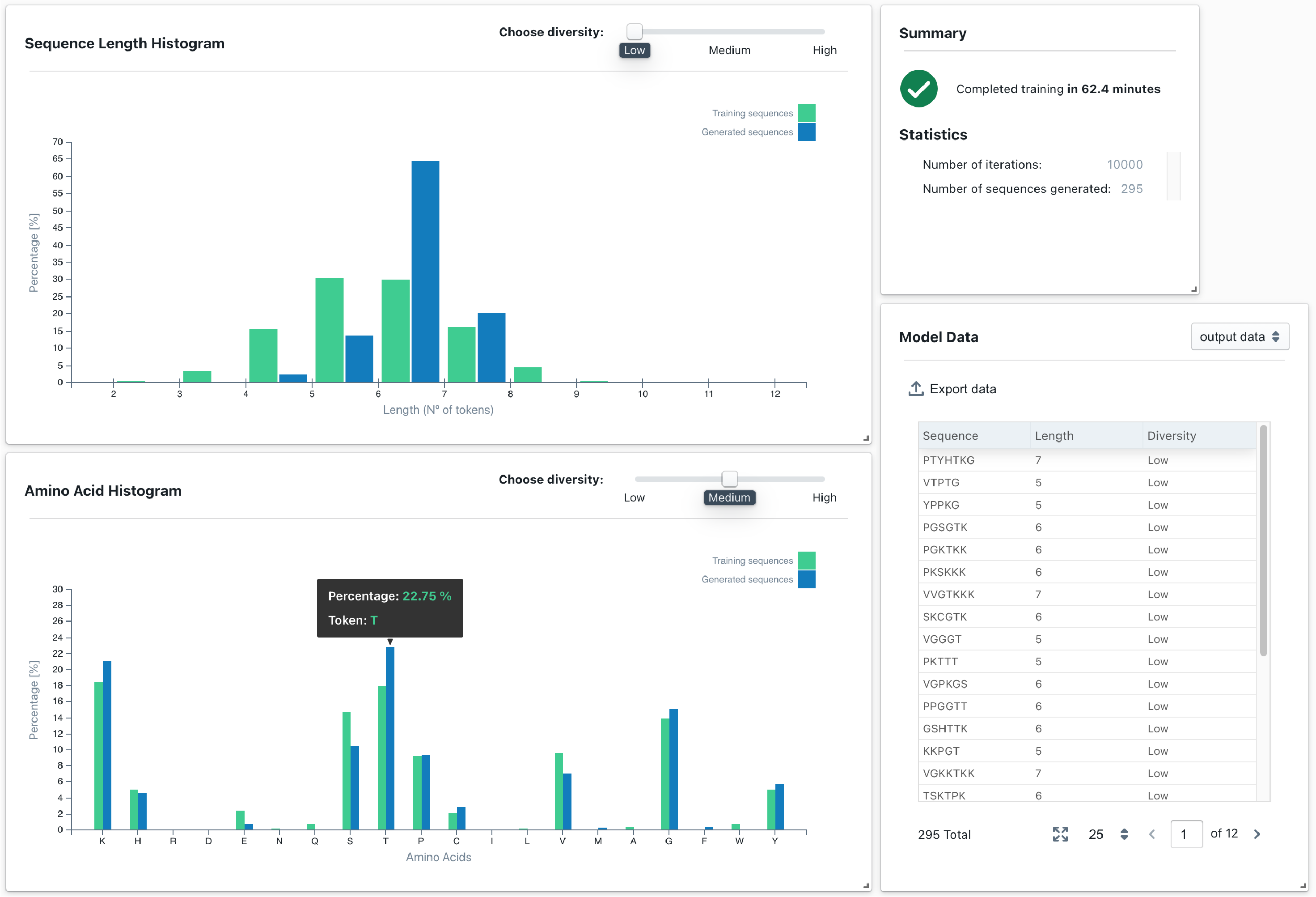
Generate new leads
Our generative models can be trained to learn the probability distribution of the molecules in a training dataset, and then used to sample it and generate new leads that follow the same distribution.
We have shown that by using some generative models we can capture the desired properties of certain peptides and protein domains.

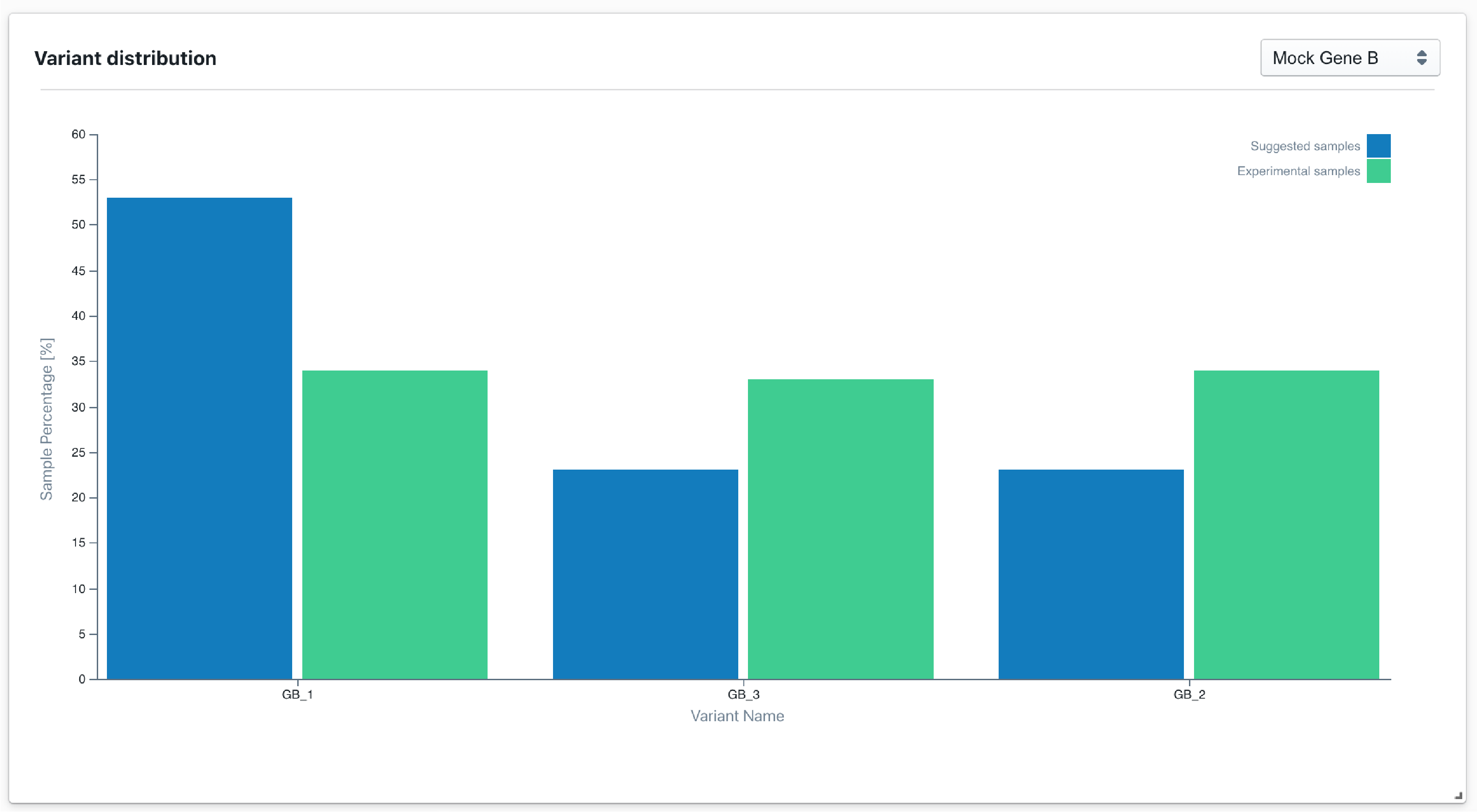
Run evolutive models
We implement bayesian optimization techniques to recommend new experiments based on empirical data gathered in the lab.
Our evolutive models can use other pre-trained predictive models as surrogate models. Evolutive models can also be used with pre-trained generative models, when exploring highly dimensional design spaces.
Deploy your own AI models directly from the Discover Module
Our platform implements advanced deep learning techniques to model biological entities, and to optimize synthetic constructs of DNA, proteins, and other biomolecules.
Train our own off-the-shelf models, or integrate our operating system with your own algorithms.